Best Practices for Single-Cell Analysis Across Modalities
Introduction
Single-cell technologies have revolutionized our understanding of cellular heterogeneity, enabling the simultaneous profiling of multiple molecular modalities. This review outlines best practices for the comprehensive analysis of single-cell data across different modalities, providing guidance on integrating and interpreting these diverse datasets.
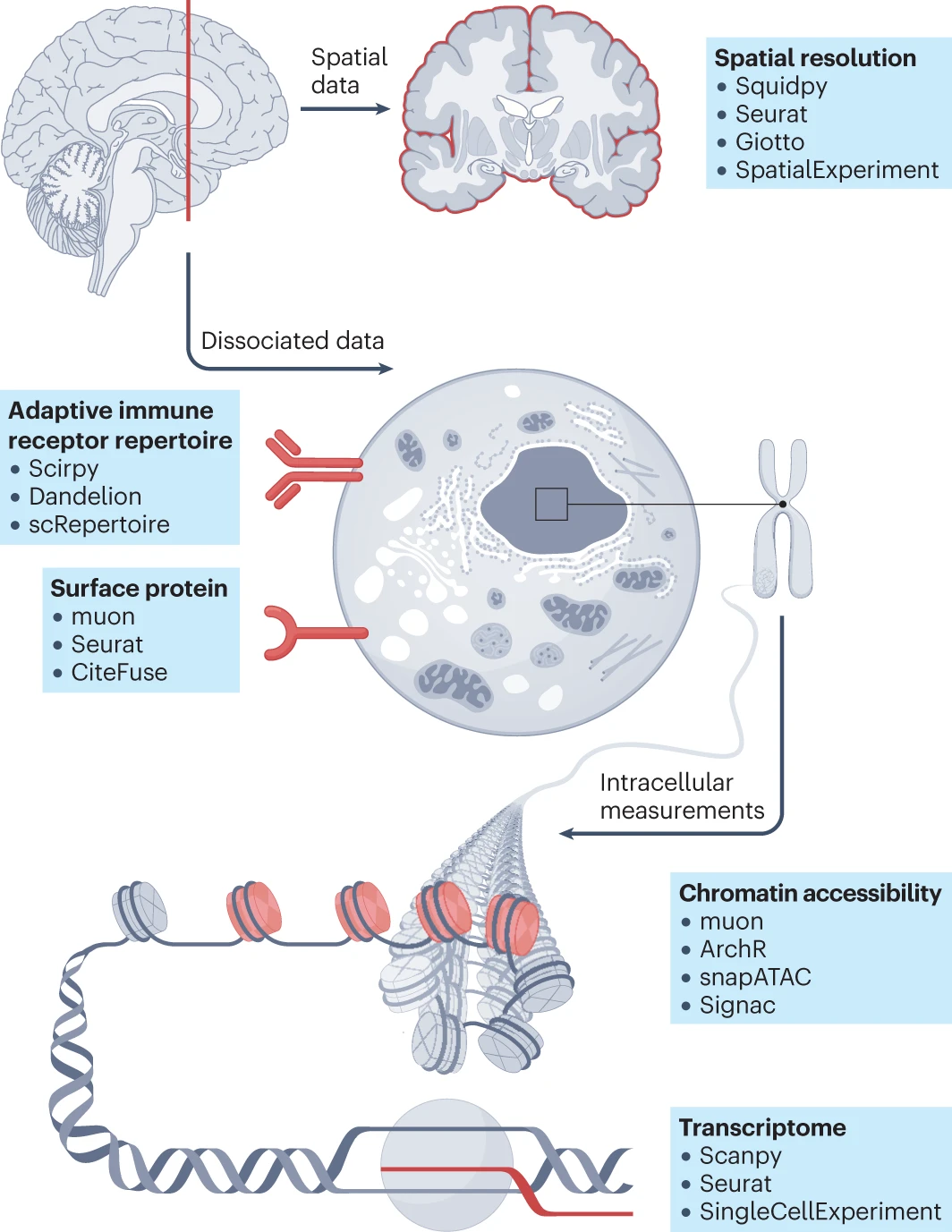
Transcriptome
scRNA-seq is the most common single-cell technique, measuring mRNA levels in individual cells. Key steps include:
- Quality Control: Filtering out low-quality cells and genes.
- Normalization: Adjusting for differences in sequencing depth.
- Clustering and Annotation: Identifying distinct cell populations and annotating them based on known markers.
- Trajectory Inference: Mapping dynamic biological processes, such as cell differentiation and development, using algorithms like Monocle and Slingshot.
Tools: Seurat, Scanpy, and Monocle.
Chromatin Accessibility
ATAC-seq (Assay for Transposase-Accessible Chromatin using sequencing) identifies regions of open chromatin, indicating active regulatory elements. Essential analysis steps include:
- Preprocessing: Quality control, alignment, and peak calling.
- Integration: Combining chromatin accessibility data with transcriptomic data to link regulatory elements with gene expression.
Tools: ArchR, snapATAC, and Cicero.
Surface Protein Expression
Techniques like CITE-seq (Cellular Indexing of Transcriptomes and Epitopes by sequencing) and REAP-seq (RNA Expression and Protein Sequencing) measure both RNA and surface protein expression in single cells. Key steps involve:
- Simultaneous Profiling: Measuring RNA and protein levels in the same cells.
- Integration: Tools like Seurat and CiteFuse enable the joint analysis of RNA and protein data, helping to correlate transcriptomic data with protein-level regulation.
Adaptive Immune Receptor Repertoires
TCR/BCR Sequencing profiles the diversity of T-cell and B-cell receptors at the single-cell level. This analysis provides insights into immune diversity and clonal expansions. Key steps include:
- Clonotype Identification: Determining unique receptor sequences.
- Repertoire Analysis: Assessing the diversity and expansion of immune cells.
Tools: Scirpy, scRepertoire, and Immunarch.
Single-cell Data Resolved in Space
Spatial Transcriptomics techniques retain spatial information about gene expression within tissues, providing context to cellular organization. Key analysis steps include:
- Spatial Mapping: Localizing gene expression data to specific regions within tissues.
- Integration: Combining spatial data with other single-cell modalities to provide a comprehensive view of cellular microenvironments.
Tools: Squidpy, SpatialExperiment, and Seurat.
Conclusions and Future Perspectives
The integration of multiple single-cell modalities is essential for a holistic understanding of cellular functions. Standardized workflows and continuous benchmarking are critical for maintaining data quality and reliability. Future advancements will likely focus on improving the resolution, throughput, and integration capabilities of single-cell technologies, leading to deeper insights into cellular biology and disease mechanisms.
For detailed insights, refer to the full article here. Best practices for single-cell analysis across modalities, Nature Review Genetics, L. Heumos et. al.

 Job
Job Collaborations
Collaborations Customer
Customer Other
Other
