Microfluidic Cell Culture: a way toward flow cells

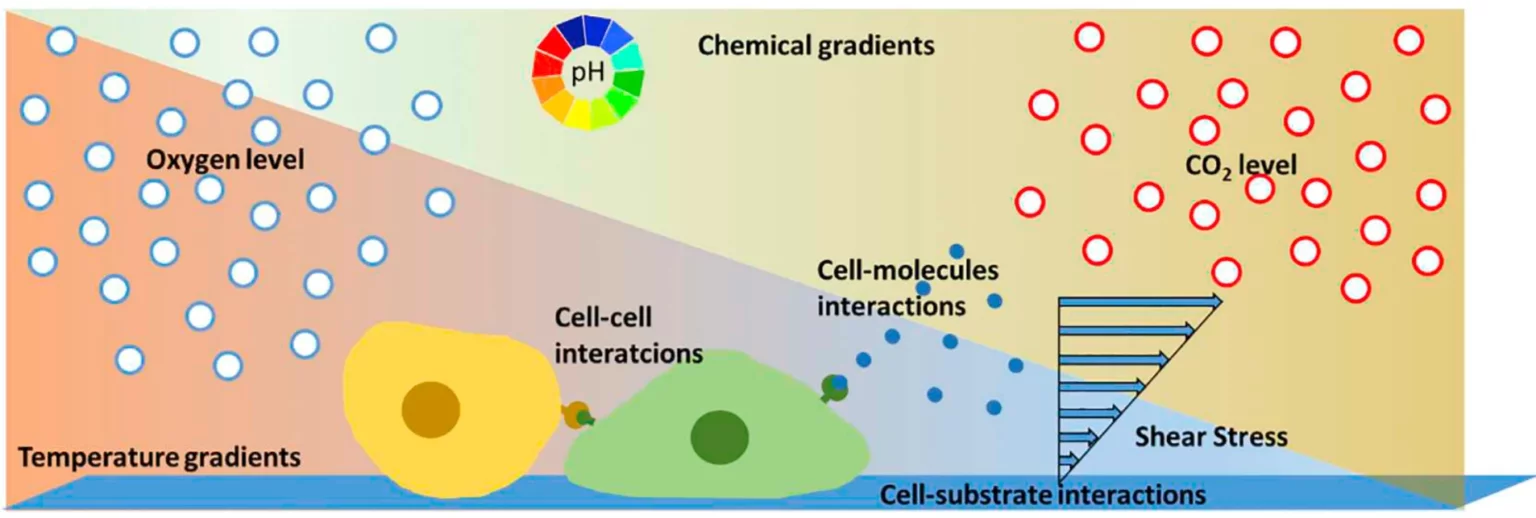
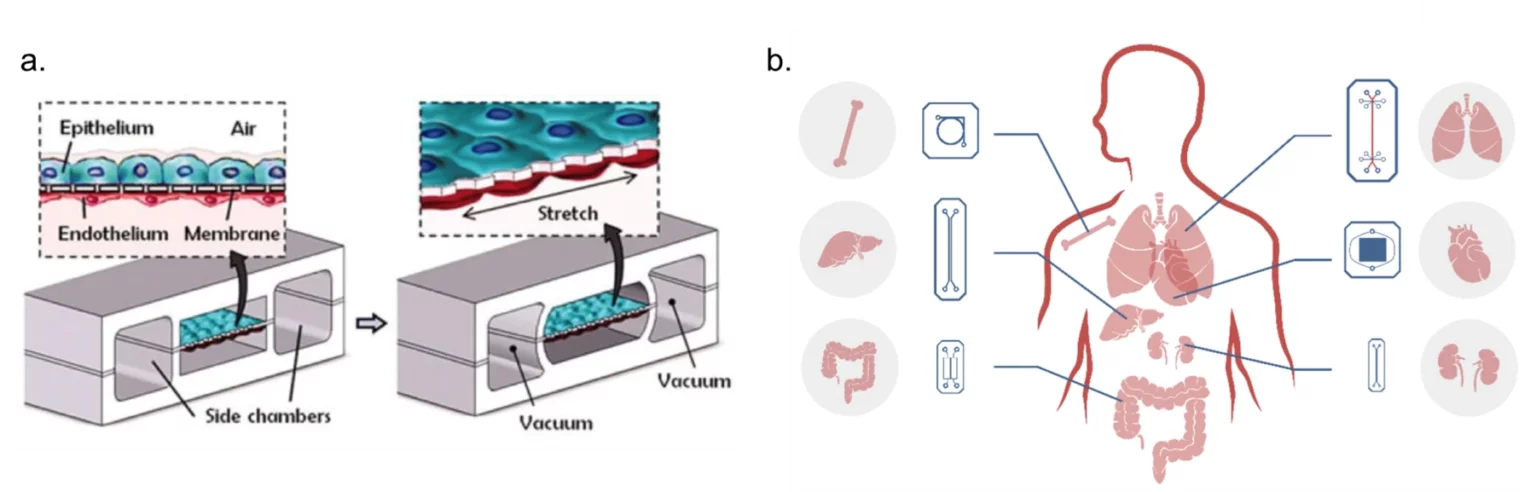
The advantages of microfluidic cell culture, which can easily mimic the cellular microenvironment and provide a high level of control in addition to automated monitoring and analysis, led to the development of the “organ-on-chip” (OoC) field in the 2010s, currently one of the most relevant areas of research in microfluidics.6 OoCs are specialised in vitro cell culture models based on microfluidic devices that are structured around three main pillars: reproducing the mechanical and biochemical stimulus of the cellular microenvironment; simulating the 3D microarchitecture of tissues with multiple cell types placed in specific communicating compartments; and, replicating functional tissue-tissue interfaces via cell-cell interaction (Fig. 2.a.). The main goal is to reproduce complex in vivo biology to better understand cellular behaviour and metabolism, disease phenotypes, and drug response without the need for insufficient in vitro models or imprecise animal models.7
Several organ-on-chip devices have been successfully developed, namely, liver8, lung7,9, kidney10, heart11,12, intestine13, bone14 and bone marrow15, nerve16, blood vessels17,18, and blood-brain barrier19 (Fig. 2.b.). They have demonstrated drug responses closer to human physiology when compared to standard in vitro models, with the enhanced complexity leading to better cell differentiation and drug transport.20 As OoC models mature, the interconnection of several OoCs in a “human-on-chip” approach becomes feasible and has been investigated by several groups in different configurations.21–24 An emerging area that has the potential to deeply change modern medicine is the development of OoC with induced pluripotent stem cells derived from patients. This has the potential to be a paradigm shift in personalised medicine and the development of patient-specific treatments.
The importance of pH monitoring
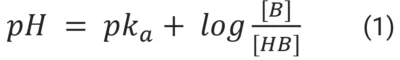
Buffering systems are capable of maintaining the pH constant due to a dynamic equilibrium. The equilibrium between the protonated and unprotonated forms of the buffer can move towards reacting to new free H+ or releasing H+ ions to compensate for ions that were added or removed from the surrounding solution. Both mechanisms have the effect of keeping the previous free H+ concentration constant to a certain extent. If the addition or removal of free H+ goes beyond a threshold that can be absorbed by the buffer, the pH will change, i.e., the buffering capacity of the solution was surpassed32. That is why complex organisms, such as mammalians, have developed dynamic buffering systems that adjust the concentration of the buffer according to the system’s needs.
The physiological buffering system is based on the equilibrium of CO2/HCO3– (bicarbonate buffering system, pKa = 6.15). The lungs, through the gas exchange of CO2, and the kidneys, through ion transport proteins, are responsible for keeping the ratio of the protonated to unprotonated species of this buffering system in homeostasis, and the pH of the organism within the physiological range (Fig. 3.a.). Cell metabolism is bound to acidify the pH of the microenvironment due to the production and release of lactate, which reacts with water to form lactic acid. For this reason, cell culture media usually contains a buffering system to keep pH under physiological conditions.33 Besides the CO2/HCO3– buffer, media can be buffered by non-volatile buffers (NVB), such as HEPES (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid) (pKa = 7.3; 37 °C), PIPES (piperazine-N,N′-bis(2-ethanesulphonic acid); pKa = 6.7) and MES (2-(N-morpholino)-ethanesulfonic acid; pKa = 6.0)34. However, NVBs in cell culture media are out of the scope of this review; detailed work on the topic can be found in Michl et al., 2019.35
The bicarbonate buffering system is replicated in cell biology labs with the aid of a CO2 incubator. This system works through the equilibrium of CO2 from the CO2-rich atmosphere of the incubator, which dissolves in the media and reacts with H2O forming carbonic acid, and the NaHCO3 present in the media (Fig. 3.b.). Cell culture media come with different concentrations of NaHCO3 and require different percentages of CO2. For example, DMEM comes with 44 mM of NaHCO3, which requires approximately 10% of CO2 in the atmosphere to maintain the pH close to 7.4. CO2 incubators are conventionally set to 5% CO2, which keeps the pH of DMEM around 7.6–7.8. The production of lactic acid and CO2 by healthy cells and the buffering capacity of serum, often added to DMEM, partially offset this difference, allowing the growth of cells with DMEM using conventional CO2 concentrations.35 However, the implications of this adjustment are not well-understood. The intracellular pH of cells is in close equilibrium with the extracellular pH of the microenvironment due to the transmembrane ion transporter proteins. Michl et al.35 studied the effect of extracellular pH change on intracellular pH and concluded that cells re-balance the internal pH based on the external one. Since most H+ targets are intracellular, this change can affect intracellular mechanisms and metabolic pathways in unpredictable ways.

When the culturing system is downscaled to micron-range in OoCs, substantially reducing the volumes, the monitoring of pH becomes even more important. Small variations in pH, e.g., 7.4 ± 0.3, are tolerated in traditional cell culture; however, the more accentuated diffusional processes of microfluidic cell culture result in a more pronounced effect on cell viability, prompting closer monitoring of crucial environmental parameters.31
Monitoring cell culture environment with flow cells
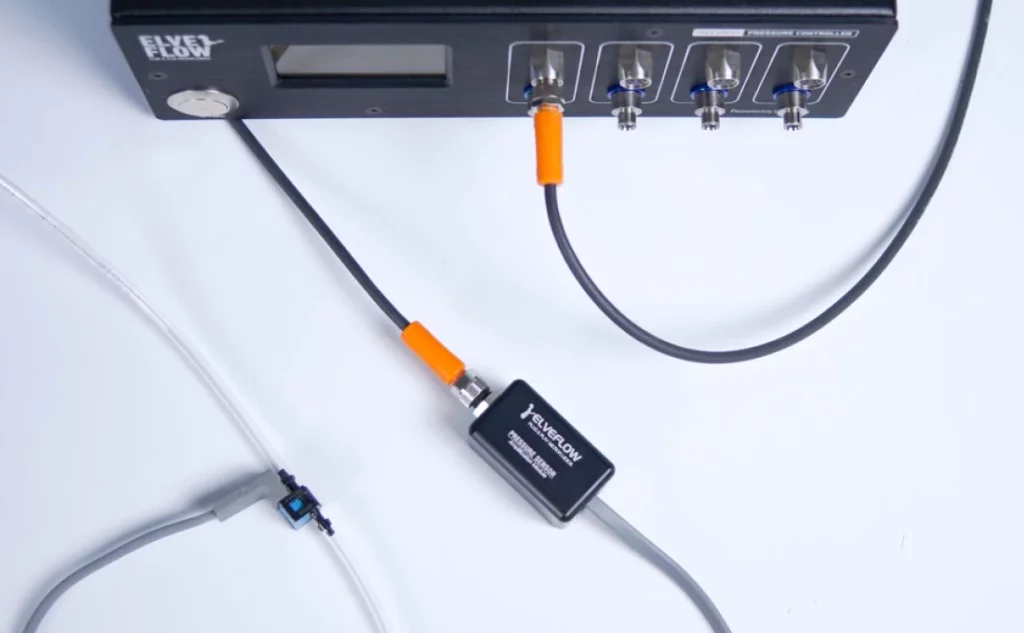
The measurement of metabolic activity in microfluidic cell culture is usually done with on-chip solutions. For example, Weltin et al.36 developed a microphysiometer system in a microfluidic chip with dissolved O2 and pH electrochemical sensors inside the culturing chamber. The authors successfully cultured brain cancer cells with online and continuous measurement of the cells’ metabolism. However, these solutions prove too complex and costly37 when the goal is to monitor the cell culture environment, which does not require the same level of detailed information. For that, microfluidic off-chip solutions, such as microfluidic flow cells, are more apt.
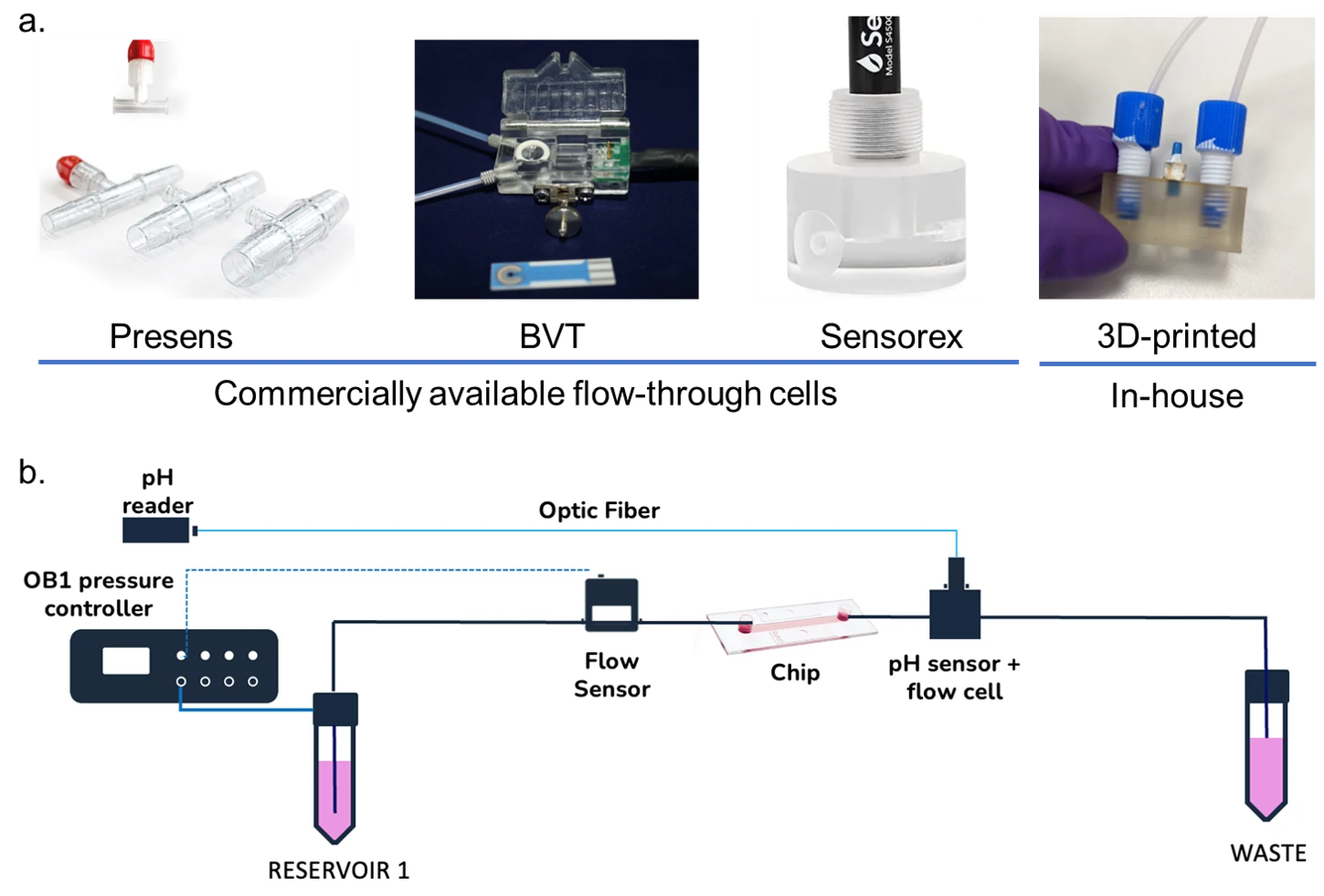
Microfluidic flow-through cells can be developed in-house38–40, which are usually 3D-printed41, or commercially available42–46. The format can vary according to the application, such as T-junctions for HPLC applications or other geometrical structures fitted to house the sensor (Fig. 4). The main goal is to place the sensors in line with the microfluidic system to allow for continuous monitoring.
A good example is the work of Zhang et al.47 who developed a “physical sensing unit” composed of pH, O2, and temperature sensors in a flow-through cell developed in-house to monitor the microenvironment of a complex organ-on-chip system. The system was designed as a modular platform to control two organ-on-chips connected to an automated flow control breadboard. Interestingly, the biomarker sensing unit, i.e., the sensors to measure the metabolism, was also off-chip. The platform was validated with a drug screening assay using liver and heart organ-on-chips, exposed to acetaminophen for 5 days and doxorubicin for 24 h, where cytotoxicity biomarkers were detected and quantified.
Farooqi et al.26 3D-printed a pH sensing flow-through cell to house an optical pH sensor built with commercially available parts that measured pH based on the colour of the phenol red present in the media. The goal was to monitor a live-on-chip system also intended to investigate the drug toxicity of doxorubicin. The researchers were able to detect a decrease in pH when cells were exposed to high concentrations of the drug when compared to the control. The same group performed similar work with a lung-on-chip system48, attesting to the robustness of the developed sensing unit. Fibroblasts had pH and O2 monitored for 3 days in a similar fashion by Ali et al.49. Also, using the phenol red of the media, Wu et al.50 designed a high-throughput pH sensing unit, which consists of parallelised microfluidic channels with a sensing chamber connected to optical pH sensors. They performed simulations to define the best shape of the sensing chamber, resulting in an oval shape, and the thickness of the PDMS layer was also decreased to minimise the loss of light transmission.
Optical sensors seem the most common choice for microfluidic flow-through cells due to their independence from reference electrodes, electrical connections, and flow rates. Also, they are not prone to biofouling, corrosion, and interference from electrochemical signals from molecules present in the medium.49
Off-chip flow cell for monitoring Conclusion and Outlook
The important growth of organ-on-chip technologies brought with it an increased need to monitor more closely what was happening inside the chips. Several on-chip and off-chip solutions have been designed, and microfluidic flow cells seem to be the way forward for microenvironment pH monitoring. On-chip sensors are suited for the detection of metabolic activity that can benefit from high spatiotemporal resolution. However, these solutions are too costly and complex for microenvironment monitoring, which does not require the same level of precision. For microenvironment monitoring, microfluidic flow cells with embedded optical sensors, which are placed in the microfluidic circuit for real-time and continuous monitoring, have been the most common choice.

Review done thanks to the support of the Protomet H2020-MSCA-ITN-2018-Action “Innovative Training Networks”, Grant agreement number: 813873.
Author: Camila Betterelli Giuliano, PhD
References
- Coluccio, M. L. et al. Microfluidic platforms for cell cultures and investigations. Microelectron. Eng. 208, 14–28 (2019).
- Lyapun, I. N., Andryukov, B. G. & Bynina, M. P. HeLa Cell Culture: Immortal Heritage of Henrietta Lacks. Mol. Genet. Microbiol. Virol. 34, 195–200 (2019).
- Mehling, M. & Tay, S. Microfluidic cell culture. Curr. Opin. Biotechnol. 25, 95–102 (2014).
- Van Norman, G. A. Limitations of Animal Studies for Predicting Toxicity in Clinical Trials: Is it Time to Rethink Our Current Approach? JACC Basic to Transl. Sci. 4, 845–854 (2019).
- Berthier, E., Surfus, J., Verbsky, J., Huttenlocher, A. & Beebe, D. An arrayed high-content chemotaxis assay for patient diagnosis. Integr. Biol. 2, 630–638 (2010).
- Convery, N. & Gadegaard, N. 30 years of microfluidics. Micro Nano Eng. 2, 76–91 (2019).
- Huh, D. et al. Reconstituting organ-level lung functions on a chip. Science (80-. ). 328, 1662–1668 (2010).
- Novik, E., Maguire, T. J., Chao, P., Cheng, K. C. & Yarmush, M. L. A microfluidic hepatic coculture platform for cell-based drug metabolism studies. Biochem. Pharmacol. 79, 1036–1044 (2010).
- Stucki, A. O. et al. A lung-on-a-chip array with an integrated bio-inspired respiration mechanism. Lab Chip 15, 1302–1310 (2015).
- Jang, K. J. et al. Human kidney proximal tubule-on-a-chip for drug transport and nephrotoxicity assessment. Integr. Biol. 5, 1119–1129 (2013).
- Grosberg, A., Alford, P. W., McCain, M. L. & Parker, K. K. Ensembles of engineered cardiac tissues for physiological and pharmacological study: Heart on a chip. Lab Chip 11, 4165–4173 (2011).
- Kobuszewska, A., Jastrzębska, E., Żukowski, K. & Brzózka, Z. Simulation of hypoxia of myocardial cells in microfluidic systems. Sci. Reports 10, 1–11 (2020).
- Kim, H. J. & Ingber, D. E. Gut-on-a-Chip microenvironment induces human intestinal cells to undergo villus differentiation. Integr. Biol. 5, 1130–1140 (2013).
- Mansoorifar, A., Gordon, R., Bergan, R. C. & Bertassoni, L. E. Bone-on-a-Chip: Microfluidic Technologies and Microphysiologic Models of Bone Tissue. Adv. Funct. Mater. 31, 2006796 (2021).
- Torisawa, Y. S. et al. Bone marrow–on–a–chip replicates hematopoietic niche physiology in vitro. Nat. Methods 11, 663–669 (2014).
- Shi, M. et al. Glia co-culture with neurons in microfluidic platforms promotes the formation and stabilization of synaptic contacts. Lab Chip 13, 3008–3021 (2013).
- Van Der Meer, A. D., Orlova, V. V., Ten Dijke, P., Van Den Berg, A. & Mummery, C. L. Three-dimensional co-cultures of human endothelial cells and embryonic stem cell-derived pericytes inside a microfluidic device. Lab Chip 13, 3562–3568 (2013).
- Yu, F., Selva Kumar, N. D., Choudhury, D., Foo, L. C. & Ng, S. H. Microfluidic platforms for modeling biological barriers in the circulatory system. Drug Discov. Today 23, 815–829 (2018).
- Booth, R. & Kim, H. Characterization of a microfluidic in vitro model of the blood-brain barrier (μBBB). Lab Chip 12, 1784–1792 (2012).
- Bhatia, S. N. & Ingber, D. E. Microfluidic organs-on-chips. Nat. Biotechnol. 32, 760–772 (2014).
- Novak, R. et al. Robotic fluidic coupling and interrogation of multiple vascularized organ chips. Nat. Biomed. Eng. 4, 407–420 (2020).
- Zhang, C., Zhao, Z., Abdul Rahim, N. A., Van Noort, D. & Yu, H. Towards a human-on-chip: Culturing multiple cell types on a chip with compartmentalized microenvironments. Lab Chip 9, 3185–3192 (2009).
- Luni, C., Serena, E. & Elvassore, N. Human-on-chip for therapy development and fundamental science. Curr. Opin. Biotechnol. 25, 45–50 (2014).
- Renggli, K., Rousset, N., Lohasz, C., Nguyen, O. T. P. & Hierlemann, A. Integrated Microphysiological Systems: Transferable Organ Models and Recirculating Flow. Adv. Biosyst. 3, 1900018 (2019).
- Schneckenburger, H. & Richter, V. Challenges in 3D live cell imaging. Photonics 8, (2021).
- Kim, B. S. et al. Real-time physiological sensor-based liver-on-chip device for monitoring drug toxicity. J. Micromechanics Microengineering 30, 115013 (2020).
- Kilic, T., Navaee, F., Stradolini, F., Renaud, P. & Carrara, S. Organs-on-chip monitoring: sensors and other strategies. Microphysiological Syst. 1, 1–1 (2018).
- Santbergen, M. J. C., van der Zande, M., Bouwmeester, H. & Nielen, M. W. F. Online and in situ analysis of organs-on-a-chip. TrAC Trends Anal. Chem. 115, 138–146 (2019).
- Begley, C. G. & Ellis, L. M. Drug development: Raise standards for preclinical cancer research. Nature 483, 531–533 (2012).
- Morita, T., Nagaki, T., Fukuda, I. & Okumura, K. Clastogenicity of low pH to various cultured mammalian cells. Mutat. Res. Mol. Mech. Mutagen. 268, 297–305 (1992).
- Lu, C. & Verbridge, S. S. Microfluidic methods for molecular biology. Microfluid. Methods Mol. Biol. 1–376 (2016). doi:10.1007/978-3-319-30019-1
- Scorpio, R. Fundamentals of Acids, Bases, Buffers & Their Application to Biochemical Systems. (Kendall Hunt Pub Co, 2000).
- Salis, A. & Monduzzi, M. Not only pH. Specific buffer effects in biological systems. Curr. Opin. Colloid Interface Sci. 23, 1–9 (2016).
- Eagle, H. Buffer combinations for mammalian cell culture. Science 174, 500–503 (1971).
- Michl, J., Park, K. C. & Swietach, P. Evidence-based guidelines for controlling pH in mammalian live-cell culture systems. Commun. Biol. 2, 1–12 (2019).
- Weltin, A. et al. Cell culture monitoring for drug screening and cancer research: a transparent, microfluidic, multi-sensor microsystem. Lab Chip 14, 138–146 (2013).
- Fuchs, S. et al. In-line analysis of organ-on-chip systems with sensors: Integration, fabrication, challenges, and potential. ACS Biomater. Sci. Eng. 7, 2926–2948 (2021).
- Pousti, M., Zarabadi, M. P., Abbaszadeh Amirdehi, M., Paquet-Mercier, F. & Greener, J. Microfluidic bioanalytical flow cells for biofilm studies: a review. Analyst 144, 68–86 (2018).
- Chauvet, A., Tibiletti, T., Caffarri, S. & Chergui, M. A microfluidic flow-cell for the study of the ultrafast dynamics of biological systems. Rev. Sci. Instrum. 85, 103118 (2014).
- Akin, M. et al. A new set up for multi-analyte sensing: At-line bio-process monitoring. Biosens. Bioelectron. 26, 4532–4537 (2011).
- Capel, A. J., Rimington, R. P., Lewis, M. P. & Christie, S. D. R. 3D printing for chemical, pharmaceutical and biological applications. Nat. Rev. Chem. 2, 422–436 (2018).
- Strebl, M. G., Bruns, M. P., Schulze, G. & Virtanen, S. Respirometric In Situ Methods for Real-Time Monitoring of Corrosion Rates: Part II. Immersion. J. Electrochem. Soc. 168, 011502 (2021).
- Ahmerkamp, S. et al. The effect of sediment grain properties and porewater flow on microbial abundance and respiration in permeable sediments. Sci. Reports 10, 1–12 (2020).
- Higuera, G. A. et al. Spatiotemporal proliferation of human stromal cells adjusts to nutrient availability and leads to stanniocalcin-1 expression in vitro and in vivo. Data Br. 5, 84–94 (2015).
- Illner, S., Hofmann, C., Löb, P. & Kragl, U. A Falling-Film Microreactor for Enzymatic Oxidation of Glucose. ChemCatChem 6, 1748–1754 (2014).
- Krejčí, J. et al. The measurement of small flow. Sensors Actuators A Phys. 266, 308–313 (2017).
- Zhang, Y. S. et al. Multisensor-integrated organs-on-chips platform for automated and continual in situ monitoring of organoid behaviors. Proc. Natl. Acad. Sci. U. S. A. 114, E2293–E2302 (2017).
- Khalid, M. A. U. et al. A lung cancer-on-chip platform with integrated biosensors for physiological monitoring and toxicity assessment. Biochem. Eng. J. 155, 107469 (2020).
- Ali, S., Shaegh, M., De Ferrari, F. & Zhang, Y. S. A microfluidic optical platform for real-time monitoring of pH and oxygen in microfluidic bioreactors and organ-on-chip devices. Biomicrofluidics 10, 044111 (2016).
- Wu, M. H., Lin, J. L., Wang, J., Cui, Z. & Cui, Z. Development of high throughput optical sensor array for on-line pH monitoring in micro-scale cell culture environment. Biomed. Microdevices 11, 265–273 (2009).
For more information or Technical discussion

Microfluidics knowledge
Do you want tips on how to best set up your microfluidic experiment? Do you need inspiration or a different angle to take on your specific problem? Well, we probably have an application note just for you, feel free to check them out!


 Job
Job Collaborations
Collaborations Customer
Customer Other
Other
